RBL4D-Var Analysis Observation Sensitivity Tutorial: Difference between revisions
m Robertson moved page PSAS Analysis Observation Sensitivity Tutorial to RBL4D-Var Analysis Observation Sensitivity Tutorial: PSAS renamed to RBL4D-Var (change visibility) |
No edit summary (change visibility) |
||
| Line 1: | Line 1: | ||
<div class="title"> | <div class="title">RBL4D-Var Observation Sensitivity</div> | ||
| Line 9: | Line 9: | ||
==Introduction== | ==Introduction== | ||
During this exercise you will apply the strong/weak constraint, dual form of 4-Dimensional Variational ('''4D-Var''') data assimilation observation sensitivity based on the | {{note}} '''Notice:''' This algorithm began based on the Physical-space Statistical Analysis System ('''PSAS''') algorithm but has evolved into a Restricted B-preconditioned Lanczos 4D-Var ([[Options#RBL4DVAR|RBL4D-Var]]). Some plots on this page still have the '''PSAS''' title but remain correct. The algorithm and ROMS [[Options|CPP Options]] relating to this data assimilation system were renamed in '''SVN revision 1022''' (May 13, 2020) and are explained in [https://www.myroms.org/projects/src/ticket/854 Trac ticket #854]. | ||
During this exercise you will apply the strong/weak constraint, dual form of 4-Dimensional Variational ('''4D-Var''') data assimilation observation sensitivity based on the Restricted B-preconditioned Lanczos 4D-Var ('''RBLanczos''') algorithm ([[Options#RBL4DVAR|RBL4D-Var]]) algorithm to ROMS configured for the U.S. west coast and the California Current System (CCS). This configuration, referred to as [[Options#WC13|WC13]], has 30 km horizontal resolution, and 30 levels in the vertical. While 30 km resolution is inadequate for capturing much of the energetic mesoscale circulation associated with the CCS, [[Options#WC13|WC13]] captures the broad-scale features of the circulation quite well, and serves as a very useful and efficient illustrative example of RBL4D-Var observation sensitivity. | |||
{{#lst:4DVar_Tutorial_Introduction|setup}} | {{#lst:4DVar_Tutorial_Introduction|setup}} | ||
==Running | ==Running RBL4D-Var Observation Sensitivity== | ||
To run this exercise, go first to the directory <span class="twilightBlue">WC13/ | To run this exercise, go first to the directory <span class="twilightBlue">WC13/RBL4DVAR_analysis_sensitivity</span>. Instructions for compiling and running the model are provided below or can be found in the <span class="twilightBlue">Readme</span> file. The recommended configuration for this exercise is one outer-loop and 50 inner-loops, and <span class="twilightBlue">roms_wc13.in</span> is configured for this default case. The number of inner-loops is controlled by the parameter [[Variables#Ninner|Ninner]] in <span class="twilightBlue">roms_wc13.in</span>. | ||
==Important CPP Options== | ==Important CPP Options== | ||
The following C-preprocessing options are activated in the [[build_Script|build script]]: | The following C-preprocessing options are activated in the [[build_Script|build script]]: | ||
<div class="box"> [[Options#RBL4DVAR_ANA_SENSITIVITY|RBL4DVAR_ANA_SENSITIVITY]] | <div class="box"> [[Options#RBL4DVAR_ANA_SENSITIVITY|RBL4DVAR_ANA_SENSITIVITY]] RBL4D-Var observation sensitivity driver<br \> [[Options#ANA_SPONGE|ANA_SPONGE]] Analytical enhanced viscosity/diffusion sponge<br /> [[Options#AD_IMPULSE|AD_IMPULSE]] Force ADM with intermittent impulses<br /> [[Options#BGQC|BGQC]] Backgound quality control of observations<br /> [[Options#MINRES|MINRES]] Minimal Residual Method for 4D-Var minimization<br /> [[Options#RPCG|RPCG]] Restricted B-preconditioned Lanczos minimization<br \> [[Options#WC13|WC13]] Application CPP option</div> | ||
==Input NetCDF Files== | ==Input NetCDF Files== | ||
| Line 25: | Line 28: | ||
==Various Scripts and Include Files== | ==Various Scripts and Include Files== | ||
The following files will be found in <span class="twilightBlue">WC13/ | The following files will be found in <span class="twilightBlue">WC13/RBL4DVAR_analysis_sensitivity</span> directory after downloading from ROMS test cases SVN repository: | ||
<div class="box"> <span class="twilightBlue">Readme</span> instructions<br /> [[build_Script|build_roms. | <div class="box"> <span class="twilightBlue">Exercise_6.pdf</span> Exercise 6 instructions<br /> <span class="twilightBlue">Readme</span> instructions<br /> [[build_Script|build_roms.csh]] csh Unix script to compile application<br /> [[build_Script|build_roms.sh]] bash shell script to compile application<br /> [[job_rbl4dvar_sen|job_rbl4dvar_sen.csh]] job configuration script<br /> [[roms.in|roms_wc13_2hours.in]] ROMS standard input script for WC13 2 hour averages<br /> [[roms.in|roms_wc13_daily.in]] ROMS standard input script for WC13 daily averages<br /> [[s4dvar.in]] 4D-Var standard input script template<br /> <span class="twilightBlue">wc13.h</span> WC13 header with CPP options</div> | ||
==Important parameters in standard input <span class="twilightBlue">roms_wc13.in</span> script== | ==Important parameters in standard input <span class="twilightBlue">roms_wc13.in</span> script== | ||
| Line 40: | Line 43: | ||
#We need to run the model application for a period that is long enough to compute meaningful circulation statistics, like mean and standard deviations for all prognostic state variables ([[Variables#zeta|zeta]], [[Variables#u|u]], [[Variables#v|v]], [[Variables#T|T]], and [[Variables#S|S]]). The standard deviations are written to NetCDF files and are read by the 4D-Var algorithm to convert modeled error correlations to error covariances. The error covariance matrix, '''D'''=diag('''B<sub>x</sub>''', '''B<sub>b</sub>''', '''B<sub>f</sub>''', '''Q'''), is very large and not well known. '''B''' is modeled as the solution of a diffusion equation as in [[Bibliography#WeaverAT_2001a|Weaver and Courtier (2001)]]. Each covariance matrix is factorized as '''B = K Σ C Σ<sup>T</sup> K<sup>T</sup>''', where '''C''' is a univariate correlation matrix, '''Σ''' is a diagonal matrix of error standard deviations, and '''K''' is a multivariate balance operator.<div class="para"> </div>In this application, we need standard deviations for initial conditions, surface forcing ([[Options#ADJUST_WSTRESS|ADJUST_WSTRESS]] and [[Options#ADJUST_STFLUX|ADJUST_STFLUX]]), and open boundary conditions ([[Options#ADJUST_BOUNDARY|ADJUST_BOUNDARY]]). If the balance operator is activated ([[Options#BALANCE_OPERATOR|BALANCE_OPERATOR]] and [[Options#ZETA_ELLIPTIC|ZETA_ELLIPTIC]]), the standard deviations for the initial and boundary conditions error covariance are in terms of the unbalanced error covariance ('''K B<sub>u</sub> K<sup>T</sup>'''). The balance operator imposes a multivariate constraint on the error covariance such that the unobserved variable information is extracted from observed data by establishing balance relationships (i.e., T-S empirical formulas, hydrostatic balance, and geostrophic balance) with other state variables ([[Bibliography#WeaverAT_2005a|Weaver ''et al.'', 2005]]). The balance operator is not used in the tutorial.<div class="para"> </div>The standard deviations for [[Options#WC13|WC13]] have already been created for you:<div class="box"><span class="twilightBlue">../Data/wc13_std_i.nc</span> initial conditions<br /><span class="twilightBlue">../Data/wc13_std_m.nc</span> model error (if weak constraint)<br /><span class="twilightBlue">../Data/wc13_std_b.nc</span> open boundary conditions<br /><span class="twilightBlue">../Data/wc13_std_f.nc</span> surface forcing (wind stress and net heat flux)</div> | #We need to run the model application for a period that is long enough to compute meaningful circulation statistics, like mean and standard deviations for all prognostic state variables ([[Variables#zeta|zeta]], [[Variables#u|u]], [[Variables#v|v]], [[Variables#T|T]], and [[Variables#S|S]]). The standard deviations are written to NetCDF files and are read by the 4D-Var algorithm to convert modeled error correlations to error covariances. The error covariance matrix, '''D'''=diag('''B<sub>x</sub>''', '''B<sub>b</sub>''', '''B<sub>f</sub>''', '''Q'''), is very large and not well known. '''B''' is modeled as the solution of a diffusion equation as in [[Bibliography#WeaverAT_2001a|Weaver and Courtier (2001)]]. Each covariance matrix is factorized as '''B = K Σ C Σ<sup>T</sup> K<sup>T</sup>''', where '''C''' is a univariate correlation matrix, '''Σ''' is a diagonal matrix of error standard deviations, and '''K''' is a multivariate balance operator.<div class="para"> </div>In this application, we need standard deviations for initial conditions, surface forcing ([[Options#ADJUST_WSTRESS|ADJUST_WSTRESS]] and [[Options#ADJUST_STFLUX|ADJUST_STFLUX]]), and open boundary conditions ([[Options#ADJUST_BOUNDARY|ADJUST_BOUNDARY]]). If the balance operator is activated ([[Options#BALANCE_OPERATOR|BALANCE_OPERATOR]] and [[Options#ZETA_ELLIPTIC|ZETA_ELLIPTIC]]), the standard deviations for the initial and boundary conditions error covariance are in terms of the unbalanced error covariance ('''K B<sub>u</sub> K<sup>T</sup>'''). The balance operator imposes a multivariate constraint on the error covariance such that the unobserved variable information is extracted from observed data by establishing balance relationships (i.e., T-S empirical formulas, hydrostatic balance, and geostrophic balance) with other state variables ([[Bibliography#WeaverAT_2005a|Weaver ''et al.'', 2005]]). The balance operator is not used in the tutorial.<div class="para"> </div>The standard deviations for [[Options#WC13|WC13]] have already been created for you:<div class="box"><span class="twilightBlue">../Data/wc13_std_i.nc</span> initial conditions<br /><span class="twilightBlue">../Data/wc13_std_m.nc</span> model error (if weak constraint)<br /><span class="twilightBlue">../Data/wc13_std_b.nc</span> open boundary conditions<br /><span class="twilightBlue">../Data/wc13_std_f.nc</span> surface forcing (wind stress and net heat flux)</div> | ||
#Since we are modeling the error covariance matrix, '''D''', we need to compute the normalization coefficients to ensure that the diagonal elements of the associated correlation matrix '''C''' are equal to unity. There are two methods to compute normalization coefficients: exact and randomization (an approximation).<div class="para"> </div>The exact method is very expensive on large grids. The normalization coefficients are computed by perturbing each model grid cell with a delta function scaled by the area (2D state variables) or volume (3D state variables), and then by convolving with the squared-root adjoint and tangent linear diffusion operators.<div class="para"> </div>The approximate method is cheaper: the normalization coefficients are computed using the randomization approach of [[Bibliography#FisherM_1995a|Fisher and Courtier (1995)]]. The coefficients are initialized with random numbers having a uniform distribution (drawn from a normal distribution with zero mean and unit variance). Then, they are scaled by the inverse squared-root of the cell area (2D state variable) or volume (3D state variable) and convolved with the squared-root adjoint and tangent diffusion operators over a specified number of iterations, Nrandom.<div class="para"> </div>Check following parameters in the 4D-Var input script [[s4dvar.in]] (see input script for details):<div class="box">[[Variables#Nmethod|Nmethod]] == 0 ! normalization method: 0=Exact (expensive) or 1=Approximated (randomization)<br />[[Variables#Nrandom|Nrandom]] == 5000 ! randomization iterations<br /><br />[[Variables#LdefNRM|LdefNRM]] == T T T T ! Create a new normalization files<br />[[Variables#LwrtNRM|LwrtNRM]] == T T T T ! Compute and write normalization<br /><br />[[Variables#CnormM|CnormM(isFsur)]] = T ! model error covariance, 2D variable at RHO-points<br />[[Variables#CnormM|CnormM(isUbar)]] = T ! model error covariance, 2D variable at U-points<br />[[Variables#CnormM|CnormM(isVbar)]] = T ! model error covariance, 2D variable at V-points<br />[[Variables#CnormM|CnormM(isUvel)]] = T ! model error covariance, 3D variable at U-points<br />[[Variables#CnormM|CnormM(isVvel)]] = T ! model error covariance, 3D variable at V-points<br />[[Variables#CnormM|CnormM(isTvar)]] = T T ! model error covariance, NT tracers<br /><br />[[Variables#CnormI|CnormI(isFsur)]] = T ! IC error covariance, 2D variable at RHO-points<br />[[Variables#CnormI|CnormI(isUbar)]] = T ! IC error covariance, 2D variable at U-points<br />[[Variables#CnormI|CnormI(isVbar)]] = T ! IC error covariance, 2D variable at V-points<br />[[Variables#CnormI|CnormI(isUvel)]] = T ! IC error covariance, 3D variable at U-points<br />[[Variables#CnormI|CnormI(isVvel)]] = T ! IC error covariance, 3D variable at V-points<br />[[Variables#CnormI|CnormI(isTvar)]] = T T ! IC error covariance, NT tracers<br /><br />[[Variables#CnormB|CnormB(isFsur)]] = T ! BC error covariance, 2D variable at RHO-points<br />[[Variables#CnormB|CnormB(isUbar)]] = T ! BC error covariance, 2D variable at U-points<br />[[Variables#CnormB|CnormB(isVbar)]] = T ! BC error covariance, 2D variable at V-points<br />[[Variables#CnormB|CnormB(isUvel)]] = T ! BC error covariance, 3D variable at U-points<br />[[Variables#CnormB|CnormB(isVvel)]] = T ! BC error covariance, 3D variable at V-points<br />[[Variables#CnormB|CnormB(isTvar)]] = T T ! BC error covariance, NT tracers<br /><br />[[Variables#CnormF|CnormF(isUstr)]] = T ! surface forcing error covariance, U-momentum stress<br />[[Variables#CnormF|CnormF(isVstr)]] = T ! surface forcing error covariance, V-momentum stress<br />[[Variables#CnormF|CnormF(isTsur)]] = T T ! surface forcing error covariance, NT tracers fluxes</div>These normalization coefficients have already been computed for you ('''../Normalization''') using the exact method since this application has a small grid (54x53x30):<div class="box"><span class="twilightBlue">../Data/wc13_nrm_i.nc</span> initial conditions<br /><span class="twilightBlue">../Data/wc13_std_m.nc</span> model error (if weak constraint)<br /><span class="twilightBlue">../Data/wc13_nrm_b.nc</span> open boundary conditions<br /><span class="twilightBlue">../Data/wc13_nrm_f.nc</span> surface forcing (wind stress and<br /> net heat flux)</div>Notice that the switches [[Variables#LdefNRM|LdefNRM]] and [[Variables#LwrtNRM|LwrtNRM]] are all '''false''' (F) since we already computed these coefficients.<div class="para"> </div>The normalization coefficients need to be computed only once for a particular application provided that the grid, land/sea masking (if any), and decorrelation scales ([[Variables#HdecayI|HdecayI]], [[Variables#VdecayI|VdecayI]], [[Variables#HdecayB|HdecayB]], [[Variables#VdecayV|VdecayV]], and [[Variables#HdecayF|HdecayF]]) remain the same. Notice that large spatial changes in the normalization coefficient structure are observed near the open boundaries and land/sea masking regions. | #Since we are modeling the error covariance matrix, '''D''', we need to compute the normalization coefficients to ensure that the diagonal elements of the associated correlation matrix '''C''' are equal to unity. There are two methods to compute normalization coefficients: exact and randomization (an approximation).<div class="para"> </div>The exact method is very expensive on large grids. The normalization coefficients are computed by perturbing each model grid cell with a delta function scaled by the area (2D state variables) or volume (3D state variables), and then by convolving with the squared-root adjoint and tangent linear diffusion operators.<div class="para"> </div>The approximate method is cheaper: the normalization coefficients are computed using the randomization approach of [[Bibliography#FisherM_1995a|Fisher and Courtier (1995)]]. The coefficients are initialized with random numbers having a uniform distribution (drawn from a normal distribution with zero mean and unit variance). Then, they are scaled by the inverse squared-root of the cell area (2D state variable) or volume (3D state variable) and convolved with the squared-root adjoint and tangent diffusion operators over a specified number of iterations, Nrandom.<div class="para"> </div>Check following parameters in the 4D-Var input script [[s4dvar.in]] (see input script for details):<div class="box">[[Variables#Nmethod|Nmethod]] == 0 ! normalization method: 0=Exact (expensive) or 1=Approximated (randomization)<br />[[Variables#Nrandom|Nrandom]] == 5000 ! randomization iterations<br /><br />[[Variables#LdefNRM|LdefNRM]] == T T T T ! Create a new normalization files<br />[[Variables#LwrtNRM|LwrtNRM]] == T T T T ! Compute and write normalization<br /><br />[[Variables#CnormM|CnormM(isFsur)]] = T ! model error covariance, 2D variable at RHO-points<br />[[Variables#CnormM|CnormM(isUbar)]] = T ! model error covariance, 2D variable at U-points<br />[[Variables#CnormM|CnormM(isVbar)]] = T ! model error covariance, 2D variable at V-points<br />[[Variables#CnormM|CnormM(isUvel)]] = T ! model error covariance, 3D variable at U-points<br />[[Variables#CnormM|CnormM(isVvel)]] = T ! model error covariance, 3D variable at V-points<br />[[Variables#CnormM|CnormM(isTvar)]] = T T ! model error covariance, NT tracers<br /><br />[[Variables#CnormI|CnormI(isFsur)]] = T ! IC error covariance, 2D variable at RHO-points<br />[[Variables#CnormI|CnormI(isUbar)]] = T ! IC error covariance, 2D variable at U-points<br />[[Variables#CnormI|CnormI(isVbar)]] = T ! IC error covariance, 2D variable at V-points<br />[[Variables#CnormI|CnormI(isUvel)]] = T ! IC error covariance, 3D variable at U-points<br />[[Variables#CnormI|CnormI(isVvel)]] = T ! IC error covariance, 3D variable at V-points<br />[[Variables#CnormI|CnormI(isTvar)]] = T T ! IC error covariance, NT tracers<br /><br />[[Variables#CnormB|CnormB(isFsur)]] = T ! BC error covariance, 2D variable at RHO-points<br />[[Variables#CnormB|CnormB(isUbar)]] = T ! BC error covariance, 2D variable at U-points<br />[[Variables#CnormB|CnormB(isVbar)]] = T ! BC error covariance, 2D variable at V-points<br />[[Variables#CnormB|CnormB(isUvel)]] = T ! BC error covariance, 3D variable at U-points<br />[[Variables#CnormB|CnormB(isVvel)]] = T ! BC error covariance, 3D variable at V-points<br />[[Variables#CnormB|CnormB(isTvar)]] = T T ! BC error covariance, NT tracers<br /><br />[[Variables#CnormF|CnormF(isUstr)]] = T ! surface forcing error covariance, U-momentum stress<br />[[Variables#CnormF|CnormF(isVstr)]] = T ! surface forcing error covariance, V-momentum stress<br />[[Variables#CnormF|CnormF(isTsur)]] = T T ! surface forcing error covariance, NT tracers fluxes</div>These normalization coefficients have already been computed for you ('''../Normalization''') using the exact method since this application has a small grid (54x53x30):<div class="box"><span class="twilightBlue">../Data/wc13_nrm_i.nc</span> initial conditions<br /><span class="twilightBlue">../Data/wc13_std_m.nc</span> model error (if weak constraint)<br /><span class="twilightBlue">../Data/wc13_nrm_b.nc</span> open boundary conditions<br /><span class="twilightBlue">../Data/wc13_nrm_f.nc</span> surface forcing (wind stress and<br /> net heat flux)</div>Notice that the switches [[Variables#LdefNRM|LdefNRM]] and [[Variables#LwrtNRM|LwrtNRM]] are all '''false''' (F) since we already computed these coefficients.<div class="para"> </div>The normalization coefficients need to be computed only once for a particular application provided that the grid, land/sea masking (if any), and decorrelation scales ([[Variables#HdecayI|HdecayI]], [[Variables#VdecayI|VdecayI]], [[Variables#HdecayB|HdecayB]], [[Variables#VdecayV|VdecayV]], and [[Variables#HdecayF|HdecayF]]) remain the same. Notice that large spatial changes in the normalization coefficient structure are observed near the open boundaries and land/sea masking regions. | ||
#Before you run this application, you need to run the standard [[ | #Before you run this application, you need to run the standard [[RBL4D-Var_Tutorial|RBL4D-Var]] ('''../RBL4DVAR''' directory) since we need the Lanczos vectors. Notice that in [[job_rbl4dvar_sen|job_rbl4dvar_sen.sh]] we have the following operation:<div class="box"><span class="red">cp -p ${Dir}/RBL4DVAR/EX3_RPCG/wc13_mod.nc wc13_lcz.nc</span></div>In 4D-Var (observartion space minimization), the Lanczos vectors are stored in the output 4D-Var NetCDF file <span class="twilightBlue">wc13_mod.nc</span>. | ||
#In addition, to run this application you need an adjoint sensitivity functional. This is computed by the following Matlab script:<div class="box"><span class="red">../Data/adsen_37N_transport.m</span></div>which creates the NetCDF file <span class="twilightBlue">wc13_ads.nc</span>. This file has already been created for you.<div class="para"> </div>The adjoint sensitivity functional is defined as the time-averaged transport crossing 37N in the upper 500m. | #In addition, to run this application you need an adjoint sensitivity functional. This is computed by the following Matlab script:<div class="box"><span class="red">../Data/adsen_37N_transport.m</span></div>which creates the NetCDF file <span class="twilightBlue">wc13_ads.nc</span>. This file has already been created for you.<div class="para"> </div>The adjoint sensitivity functional is defined as the time-averaged transport crossing 37N in the upper 500m. | ||
#Customize your preferred [[build_Script|build script]] and provide the appropriate values for: | #Customize your preferred [[build_Script|build script]] and provide the appropriate values for: | ||
| Line 47: | Line 50: | ||
#*Fortran compiler, <span class="salmon">FORT</span> | #*Fortran compiler, <span class="salmon">FORT</span> | ||
#*MPI flags, <span class="salmon">USE_MPI</span> and <span class="salmon">USE_MPIF90</span> | #*MPI flags, <span class="salmon">USE_MPI</span> and <span class="salmon">USE_MPIF90</span> | ||
#*Path of MPI, NetCDF, and ARPACK libraries according to the compiler are set in [[my_build_paths. | #*Path of MPI, NetCDF, and ARPACK libraries according to the compiler are set in [[build_Script#Library_and_Executable_Paths|my_build_paths.csh]]. Notice that you need to provide the correct places of these libraries for your computer. If you want to ignore this section, set <span class="salmon">USE_MY_LIBS</span> value to '''no'''. | ||
#Notice that the most important CPP options for this application are specified in the [[build_Script|build script]] instead of <span class="twilightBlue">wc13.h</span>:<div class="box"><span class="twilightBlue">setenv MY_CPP_FLAGS "${MY_CPP_FLAGS} -DRBL4DVAR_ANA_SENSITIVITY"<br />setenv MY_CPP_FLAGS "${MY_CPP_FLAGS} -DAD_IMPULSE"<br />setenv MY_CPP_FLAGS "${MY_CPP_FLAGS} -DRPCG"</span></div>This is to allow flexibility with different CPP options.<div class="para"> </div>For this to work, however, any '''#undef''' directives MUST be avoided in the header file <span class="twilightBlue">wc13.h</span> since it has precedence during C-preprocessing. | #Notice that the most important CPP options for this application are specified in the [[build_Script|build script]] instead of <span class="twilightBlue">wc13.h</span>:<div class="box"><span class="twilightBlue">setenv MY_CPP_FLAGS "${MY_CPP_FLAGS} -DRBL4DVAR_ANA_SENSITIVITY"<br />setenv MY_CPP_FLAGS "${MY_CPP_FLAGS} -DAD_IMPULSE"<br />setenv MY_CPP_FLAGS "${MY_CPP_FLAGS} -DRPCG"</span></div>This is to allow flexibility with different CPP options.<div class="para"> </div>For this to work, however, any '''#undef''' directives MUST be avoided in the header file <span class="twilightBlue">wc13.h</span> since it has precedence during C-preprocessing. | ||
#You MUST use the [[build_Script|build script]] to compile. | #You MUST use the [[build_Script|build script]] to compile. | ||
#Customize the ROMS input script <span class="twilightBlue">roms_wc13.in</span> and specify the appropriate values for the distributed-memory partition. It is set by default to:<div class="box">[[Variables#NtileI|NtileI]] == 2 ! I-direction partition<br />[[Variables#NtileJ|NtileJ]] == 4 ! J-direction partition</div>Notice that the adjoint-based algorithms can only be run in parallel using MPI. This is because of the way that the adjoint model is constructed. | #Customize the ROMS input script <span class="twilightBlue">roms_wc13.in</span> and specify the appropriate values for the distributed-memory partition. It is set by default to:<div class="box">[[Variables#NtileI|NtileI]] == 2 ! I-direction partition<br />[[Variables#NtileJ|NtileJ]] == 4 ! J-direction partition</div>Notice that the adjoint-based algorithms can only be run in parallel using MPI. This is because of the way that the adjoint model is constructed. | ||
#Customize the configuration script [[ | #Customize the configuration script [[job_rbl4dvar_sen|job_rbl4dvar_sen.sh]] and provide the appropriate place for the [[substitute]] Perl script:<div class="box"><span class="twilightBlue">set SUBSTITUTE=${ROMS_ROOT}/ROMS/Bin/substitute</span></div>This script is distributed with ROMS and it is found in the ROMS/Bin sub-directory. Alternatively, you can define ROMS_ROOT environmental variable in your .cshrc login script. For example, I have:<div class="box"><span class="twilightBlue">setenv ROMS_ROOT /home/arango/ocean/toms/repository/trunk</span></div> | ||
#Execute the configuration [[ | #Execute the configuration [[job_rbl4dvar_sen|job_rbl4dvar_sen.sh]] '''before''' running the model. It copies the required files and creates <span class="twilightBlue">rbl4dvar.in</span> input script from template '''[[s4dvar.in]]'''. This has to be done '''every time''' that you run this application. We need a clean and fresh copy of the initial conditions and observation files since they are modified by ROMS during execution. | ||
#Run ROMS with data assimilation:<div class="box"><span class="red">mpirun -np 8 romsM roms_wc13.in > & log &</span></div> | #Run ROMS with data assimilation:<div class="box"><span class="red">mpirun -np 8 romsM roms_wc13.in > & log &</span></div> | ||
#We recommend creating a new subdirectory <span class="twilightBlue">EX6</span>, and saving the solution in it for analysis and plotting to avoid overwriting solutions when playing with different parameters. For example<div class="box">mkdir EX6<br />mv Build_roms | #We recommend creating a new subdirectory <span class="twilightBlue">EX6</span>, and saving the solution in it for analysis and plotting to avoid overwriting solutions when playing with different parameters. For example<div class="box">mkdir EX6<br />mv Build_roms rbl4dvar.in *.nc log EX6<br />cp -p romsM roms_wc13.in EX6</div>where log is the ROMS standard output specified in the previous step. | ||
==Results== | ==Results== | ||
The <span class="twilightBlue">WC13/plotting/ | The <span class="twilightBlue">WC13/plotting/plot_rbl4dvar_analysis_sensitivity.m</span> Matlab script will allow you to plot the RBL4D-Var observation sensitivity: | ||
{|align="center" | {|align="center" | ||
|- | |- | ||
|[[Image:psas_sensitivity_2019.png|500px|thumb|center|<center> | |[[Image:psas_sensitivity_2019.png|500px|thumb|center|<center>RBL4D-Var Analysis Observation Sensitivity<br />''prior'' saved daily</center>]] | ||
|[[Image:psas_sensitivity_2hour_2019.png|500px|thumb|center|<center> | |[[Image:psas_sensitivity_2hour_2019.png|500px|thumb|center|<center>RBL4D-Var Analysis Observation Sensitivity<br />''prior'' saved every 2 hours</center>]] | ||
|} | |} | ||
<div style="clear: both;"></div> | <div style="clear: both;"></div> | ||
Revision as of 18:23, 24 July 2020
Introduction
![]() Notice: This algorithm began based on the Physical-space Statistical Analysis System (PSAS) algorithm but has evolved into a Restricted B-preconditioned Lanczos 4D-Var (RBL4D-Var). Some plots on this page still have the PSAS title but remain correct. The algorithm and ROMS CPP Options relating to this data assimilation system were renamed in SVN revision 1022 (May 13, 2020) and are explained in Trac ticket #854.
Notice: This algorithm began based on the Physical-space Statistical Analysis System (PSAS) algorithm but has evolved into a Restricted B-preconditioned Lanczos 4D-Var (RBL4D-Var). Some plots on this page still have the PSAS title but remain correct. The algorithm and ROMS CPP Options relating to this data assimilation system were renamed in SVN revision 1022 (May 13, 2020) and are explained in Trac ticket #854.
During this exercise you will apply the strong/weak constraint, dual form of 4-Dimensional Variational (4D-Var) data assimilation observation sensitivity based on the Restricted B-preconditioned Lanczos 4D-Var (RBLanczos) algorithm (RBL4D-Var) algorithm to ROMS configured for the U.S. west coast and the California Current System (CCS). This configuration, referred to as WC13, has 30 km horizontal resolution, and 30 levels in the vertical. While 30 km resolution is inadequate for capturing much of the energetic mesoscale circulation associated with the CCS, WC13 captures the broad-scale features of the circulation quite well, and serves as a very useful and efficient illustrative example of RBL4D-Var observation sensitivity.
Model Set-up
The WC13 model domain is shown in Fig. 1 and has open boundaries along the northern, western, and southern edges of the model domain.
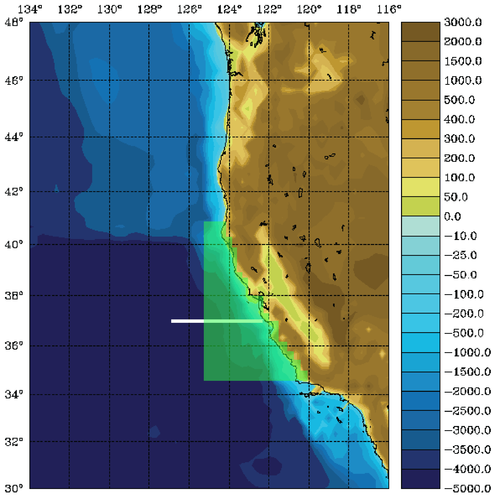
In the tutorial, you will perform a 4D-Var data assimilation cycle that spans the period 3-6 January, 2004. The 4D-Var control vector δz is comprised of increments to the initial conditions, δx(t0), surface forcing, δf(t), and open boundary conditions, δb(t). The prior initial conditions, xb(t0), are taken from the sequence of 4D-Var experiments described by Moore et al. (2011b) in which data were assimilated every 7 days during the period July 2002- December 2004. The prior surface forcing, fb(t), takes the form of surface wind stress, heat flux, and a freshwater flux computed using the ROMS bulk flux formulation, and using near surface air data from COAMPS (Doyle et al., 2009). Clamped open boundary conditions are imposed on (u,v) and tracers, and the prior boundary conditions, bb(t), are taken from the global ECCO product (Wunsch and Heimbach, 2007). The free-surface height and vertically integrated velocity components are subject to the usual Chapman and Flather radiation conditions at the open boundaries. The prior surface forcing and open boundary conditions are provided daily and linearly interpolated in time. Similarly, the increments δf(t) and δb(t) are also computed daily and linearly interpolated in time.
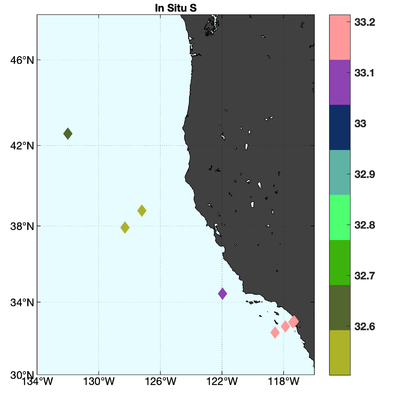
The observations assimilated into the model are satellite SST, satellite SSH in the form of a gridded product from Aviso, and hydrographic observations of temperature and salinity collected from Argo floats and during the GLOBEC/LTOP and CalCOFI cruises off the coast of Oregon and southern California, respectively. The observation locations are illustrated in Fig. 2.
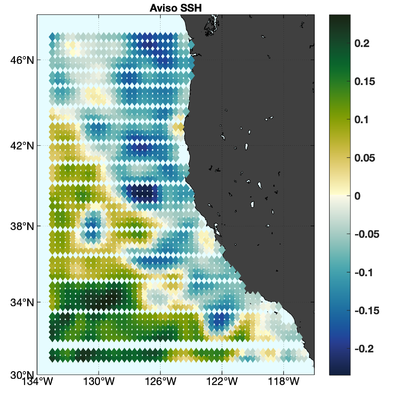 |
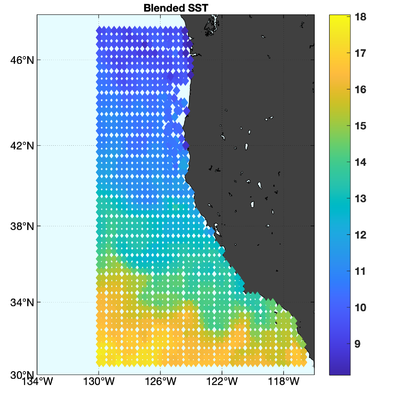 |
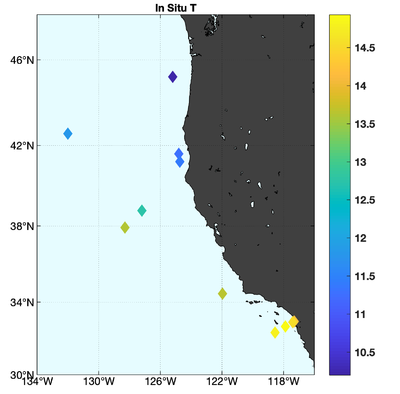 |
 |
Running RBL4D-Var Observation Sensitivity
To run this exercise, go first to the directory WC13/RBL4DVAR_analysis_sensitivity. Instructions for compiling and running the model are provided below or can be found in the Readme file. The recommended configuration for this exercise is one outer-loop and 50 inner-loops, and roms_wc13.in is configured for this default case. The number of inner-loops is controlled by the parameter Ninner in roms_wc13.in.
Important CPP Options
The following C-preprocessing options are activated in the build script:
ANA_SPONGE Analytical enhanced viscosity/diffusion sponge
AD_IMPULSE Force ADM with intermittent impulses
BGQC Backgound quality control of observations
MINRES Minimal Residual Method for 4D-Var minimization
RPCG Restricted B-preconditioned Lanczos minimization
WC13 Application CPP option
Input NetCDF Files
WC13 requires the following input NetCDF files:
Nonlinear Initial File: wc13_ini.nc
Forcing File 01: ../Data/coamps_wc13_lwrad_down.nc
Forcing File 02: ../Data/coamps_wc13_Pair.nc
Forcing File 03: ../Data/coamps_wc13_Qair.nc
Forcing File 04: ../Data/coamps_wc13_rain.nc
Forcing File 05: ../Data/coamps_wc13_swrad.nc
Forcing File 06: ../Data/coamps_wc13_Tair.nc
Forcing File 07: ../Data/coamps_wc13_wind.nc
Boundary File: ../Data/wc13_ecco_bry.nc
Adjoint Sensitivity File: wc13_ads.nc
Initial Conditions STD File: ../Data/wc13_std_i.nc
Model STD File: ../Data/wc13_std_m.nc
Boundary Conditions STD File: ../Data/wc13_std_b.nc
Surface Forcing STD File: ../Data/wc13_std_f.nc
Initial Conditions Norm File: ../Data/wc13_nrm_i.nc
Model Norm File: ../Data/wc13_nrm_m.nc
Boundary Conditions Norm File: ../Data/wc13_nrm_b.nc
Surface Forcing Norm File: ../Data/wc13_nrm_f.nc
Observations File: wc13_obs.nc
Lanczos Vectors File: wc13_lcz.nc
Various Scripts and Include Files
The following files will be found in WC13/RBL4DVAR_analysis_sensitivity directory after downloading from ROMS test cases SVN repository:
Readme instructions
build_roms.csh csh Unix script to compile application
build_roms.sh bash shell script to compile application
job_rbl4dvar_sen.csh job configuration script
roms_wc13_2hours.in ROMS standard input script for WC13 2 hour averages
roms_wc13_daily.in ROMS standard input script for WC13 daily averages
s4dvar.in 4D-Var standard input script template
wc13.h WC13 header with CPP options
Important parameters in standard input roms_wc13.in script
- Notice that this driver uses the following adjoint sensitivity parameters (see input script for details):
- DstrS == 0.0d0 ! starting day
DendS == 0.0d0 ! ending day
KstrS == 1 ! starting level
KendS == 30 ! ending level
Lstate(isFsur) == T ! free-surface
Lstate(isUbar) == T ! 2D U-momentum
Lstate(isVbar) == T ! 2D V-momentum
Lstate(isUvel) == T ! 3D U-momentum
Lstate(isVvel) == T ! 3D V-momentum
Lstate(isWvel) == F ! 3D W-momentum
Lstate(isTvar) == T T ! tracers
- Both FWDNAME and HISNAME must be the same:
Instructions
To run this application you need to take the following steps:
- We need to run the model application for a period that is long enough to compute meaningful circulation statistics, like mean and standard deviations for all prognostic state variables (zeta, u, v, T, and S). The standard deviations are written to NetCDF files and are read by the 4D-Var algorithm to convert modeled error correlations to error covariances. The error covariance matrix, D=diag(Bx, Bb, Bf, Q), is very large and not well known. B is modeled as the solution of a diffusion equation as in Weaver and Courtier (2001). Each covariance matrix is factorized as B = K Σ C ΣT KT, where C is a univariate correlation matrix, Σ is a diagonal matrix of error standard deviations, and K is a multivariate balance operator.In this application, we need standard deviations for initial conditions, surface forcing (ADJUST_WSTRESS and ADJUST_STFLUX), and open boundary conditions (ADJUST_BOUNDARY). If the balance operator is activated (BALANCE_OPERATOR and ZETA_ELLIPTIC), the standard deviations for the initial and boundary conditions error covariance are in terms of the unbalanced error covariance (K Bu KT). The balance operator imposes a multivariate constraint on the error covariance such that the unobserved variable information is extracted from observed data by establishing balance relationships (i.e., T-S empirical formulas, hydrostatic balance, and geostrophic balance) with other state variables (Weaver et al., 2005). The balance operator is not used in the tutorial.The standard deviations for WC13 have already been created for you:../Data/wc13_std_i.nc initial conditions
../Data/wc13_std_m.nc model error (if weak constraint)
../Data/wc13_std_b.nc open boundary conditions
../Data/wc13_std_f.nc surface forcing (wind stress and net heat flux) - Since we are modeling the error covariance matrix, D, we need to compute the normalization coefficients to ensure that the diagonal elements of the associated correlation matrix C are equal to unity. There are two methods to compute normalization coefficients: exact and randomization (an approximation).The exact method is very expensive on large grids. The normalization coefficients are computed by perturbing each model grid cell with a delta function scaled by the area (2D state variables) or volume (3D state variables), and then by convolving with the squared-root adjoint and tangent linear diffusion operators.The approximate method is cheaper: the normalization coefficients are computed using the randomization approach of Fisher and Courtier (1995). The coefficients are initialized with random numbers having a uniform distribution (drawn from a normal distribution with zero mean and unit variance). Then, they are scaled by the inverse squared-root of the cell area (2D state variable) or volume (3D state variable) and convolved with the squared-root adjoint and tangent diffusion operators over a specified number of iterations, Nrandom.Check following parameters in the 4D-Var input script s4dvar.in (see input script for details):Nmethod == 0 ! normalization method: 0=Exact (expensive) or 1=Approximated (randomization)These normalization coefficients have already been computed for you (../Normalization) using the exact method since this application has a small grid (54x53x30):
Nrandom == 5000 ! randomization iterations
LdefNRM == T T T T ! Create a new normalization files
LwrtNRM == T T T T ! Compute and write normalization
CnormM(isFsur) = T ! model error covariance, 2D variable at RHO-points
CnormM(isUbar) = T ! model error covariance, 2D variable at U-points
CnormM(isVbar) = T ! model error covariance, 2D variable at V-points
CnormM(isUvel) = T ! model error covariance, 3D variable at U-points
CnormM(isVvel) = T ! model error covariance, 3D variable at V-points
CnormM(isTvar) = T T ! model error covariance, NT tracers
CnormI(isFsur) = T ! IC error covariance, 2D variable at RHO-points
CnormI(isUbar) = T ! IC error covariance, 2D variable at U-points
CnormI(isVbar) = T ! IC error covariance, 2D variable at V-points
CnormI(isUvel) = T ! IC error covariance, 3D variable at U-points
CnormI(isVvel) = T ! IC error covariance, 3D variable at V-points
CnormI(isTvar) = T T ! IC error covariance, NT tracers
CnormB(isFsur) = T ! BC error covariance, 2D variable at RHO-points
CnormB(isUbar) = T ! BC error covariance, 2D variable at U-points
CnormB(isVbar) = T ! BC error covariance, 2D variable at V-points
CnormB(isUvel) = T ! BC error covariance, 3D variable at U-points
CnormB(isVvel) = T ! BC error covariance, 3D variable at V-points
CnormB(isTvar) = T T ! BC error covariance, NT tracers
CnormF(isUstr) = T ! surface forcing error covariance, U-momentum stress
CnormF(isVstr) = T ! surface forcing error covariance, V-momentum stress
CnormF(isTsur) = T T ! surface forcing error covariance, NT tracers fluxes../Data/wc13_nrm_i.nc initial conditionsNotice that the switches LdefNRM and LwrtNRM are all false (F) since we already computed these coefficients.
../Data/wc13_std_m.nc model error (if weak constraint)
../Data/wc13_nrm_b.nc open boundary conditions
../Data/wc13_nrm_f.nc surface forcing (wind stress and
net heat flux)The normalization coefficients need to be computed only once for a particular application provided that the grid, land/sea masking (if any), and decorrelation scales (HdecayI, VdecayI, HdecayB, VdecayV, and HdecayF) remain the same. Notice that large spatial changes in the normalization coefficient structure are observed near the open boundaries and land/sea masking regions. - Before you run this application, you need to run the standard RBL4D-Var (../RBL4DVAR directory) since we need the Lanczos vectors. Notice that in job_rbl4dvar_sen.sh we have the following operation:cp -p ${Dir}/RBL4DVAR/EX3_RPCG/wc13_mod.nc wc13_lcz.ncIn 4D-Var (observartion space minimization), the Lanczos vectors are stored in the output 4D-Var NetCDF file wc13_mod.nc.
- In addition, to run this application you need an adjoint sensitivity functional. This is computed by the following Matlab script:../Data/adsen_37N_transport.mwhich creates the NetCDF file wc13_ads.nc. This file has already been created for you.The adjoint sensitivity functional is defined as the time-averaged transport crossing 37N in the upper 500m.
- Customize your preferred build script and provide the appropriate values for:
- Root directory, MY_ROOT_DIR
- ROMS source code, MY_ROMS_SRC
- Fortran compiler, FORT
- MPI flags, USE_MPI and USE_MPIF90
- Path of MPI, NetCDF, and ARPACK libraries according to the compiler are set in my_build_paths.csh. Notice that you need to provide the correct places of these libraries for your computer. If you want to ignore this section, set USE_MY_LIBS value to no.
- Notice that the most important CPP options for this application are specified in the build script instead of wc13.h:setenv MY_CPP_FLAGS "${MY_CPP_FLAGS} -DRBL4DVAR_ANA_SENSITIVITY"This is to allow flexibility with different CPP options.
setenv MY_CPP_FLAGS "${MY_CPP_FLAGS} -DAD_IMPULSE"
setenv MY_CPP_FLAGS "${MY_CPP_FLAGS} -DRPCG"For this to work, however, any #undef directives MUST be avoided in the header file wc13.h since it has precedence during C-preprocessing. - You MUST use the build script to compile.
- Customize the ROMS input script roms_wc13.in and specify the appropriate values for the distributed-memory partition. It is set by default to:Notice that the adjoint-based algorithms can only be run in parallel using MPI. This is because of the way that the adjoint model is constructed.
- Customize the configuration script job_rbl4dvar_sen.sh and provide the appropriate place for the substitute Perl script:set SUBSTITUTE=${ROMS_ROOT}/ROMS/Bin/substituteThis script is distributed with ROMS and it is found in the ROMS/Bin sub-directory. Alternatively, you can define ROMS_ROOT environmental variable in your .cshrc login script. For example, I have:setenv ROMS_ROOT /home/arango/ocean/toms/repository/trunk
- Execute the configuration job_rbl4dvar_sen.sh before running the model. It copies the required files and creates rbl4dvar.in input script from template s4dvar.in. This has to be done every time that you run this application. We need a clean and fresh copy of the initial conditions and observation files since they are modified by ROMS during execution.
- Run ROMS with data assimilation:mpirun -np 8 romsM roms_wc13.in > & log &
- We recommend creating a new subdirectory EX6, and saving the solution in it for analysis and plotting to avoid overwriting solutions when playing with different parameters. For examplemkdir EX6where log is the ROMS standard output specified in the previous step.
mv Build_roms rbl4dvar.in *.nc log EX6
cp -p romsM roms_wc13.in EX6
Results
The WC13/plotting/plot_rbl4dvar_analysis_sensitivity.m Matlab script will allow you to plot the RBL4D-Var observation sensitivity:
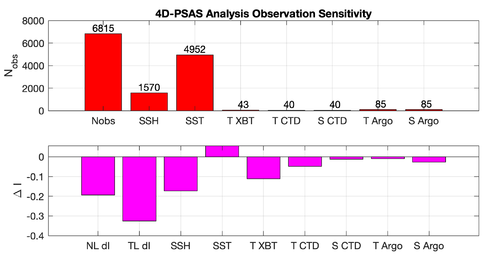 prior saved daily |
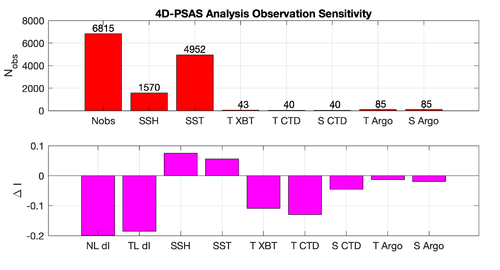 prior saved every 2 hours |