easygrid
EASYGRID is a simple matlab script to make ROMS grids and initialization files. It is an austere grid-making alternative designed as a beginner's tool, yet capable of making grids for realistic applications. Since most of EASYGRID is in one file (easygrid.m), it is easier to follow and understand the steps required to generate a simple grid (i.e. useful as a learning or teaching tool) and also reduces the required steps to get the software up and running (i.e. downloading and compiling dependencies).
This WikiROMS page is mainly a tutorial, where I provide test bathymetry and coastline files so you can experiment with EASYGRID. Also, this page is where I will post news and updates.
| QUICK LINKS: Hosting page at Google-Code Discussion Forum |
NEWS: 24 April 2008: Version 0 is out |
EASYGRID v0 is in BETA stage and could have problems. Report HERE not only bugs, but also report successful use. Once the bug:success ratio decreases... I will remove this sign. |
Download EASYGRID
- Download EASYGRID (This download also includes the bathymetry and coastline files required for this tutorial)
- Extract (i.e. unzip or unpack) the file in a location where it can stay indefinitely.
- Add the location where you placed EASYGRID to your Matlab search path (see #5 as an example).
- Dependencies: EASYGRID uses MEXNC, SNCTOOLS and ROMS Matlab tool-kit (only a few scripts like spheriq_dist.m , smth_bath.m , pcolorjw.m).
 For a tutorial on how to download/install these dependencies, CLICK HERE.
For a tutorial on how to download/install these dependencies, CLICK HERE.
Getting bathymetry
EASYGRID needs a bathymetry .mat file containing 3 vectors (xbathy, ybathy and zbathy), where:
- xbathy = longitude
- ybathy = latitude
- zbathy = depth (positive, in meters... For more information about bathymetry for ROMS, click HERE)
 NOTE: Vectors xbathy and ybathy are in decimal degrees.
NOTE: Vectors xbathy and ybathy are in decimal degrees.
There easiest way to get bathymetry data (that I know) is to use Rich Signell's read_srtm30plus.m Matlab script. Instant bathymetry in 1 line of Matlab code! Click HERE for instructions to get and use read_srtm30plus.m.
The most difficult way to get bathymetry data (that I know) is to scan a chart and then digitize it using your mouse and a digitizing software like Didger (sorry, I am not aware of any good digitizing free software).
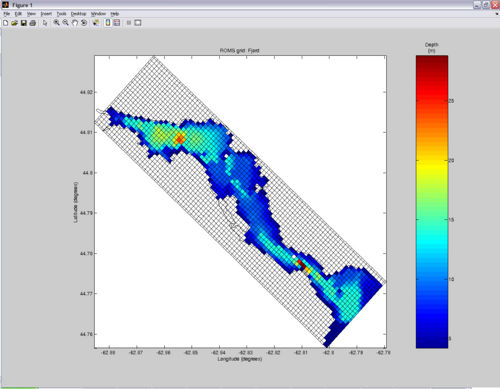
For this tutorial, you will need Fjord_bathy.mat. You probably already have this file since it is included in EASYGRID_v0.rar (file from the "Download" section). I created Fjord_bathy.mat by merging two bathymetry files... one high-resolution digitized from a chart and another coarser-resolution using read_srtm30plus.m (see Figure).
![]() If you want to reproduce this plot as an exercise, copy-paste the following code on Matlab's Command Window:
If you want to reproduce this plot as an exercise, copy-paste the following code on Matlab's Command Window:
load Fjord_bathy.mat plot(xbathy,ybathy,'.'); xlabel('Longitude'); ylabel('Latitude'); axis square;
Getting coastline
In EASYGRID, the coastline is only used for plotting and is therefore not indispensable. However, it is handy to have a coastline for reference, especially when editing the land/sea mask. For EASYGRID, the coastline should be 2 vectors named "lat" and "lon" (both in units of decimal degrees) contained in a .mat file.
One way to get coastline data is at GSHHG. NOAA's Coastline Extractor is no longer available.
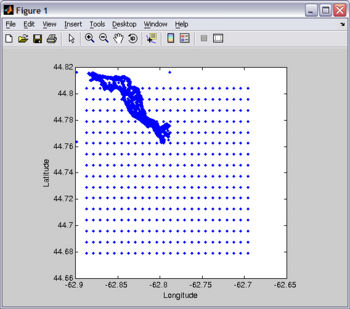
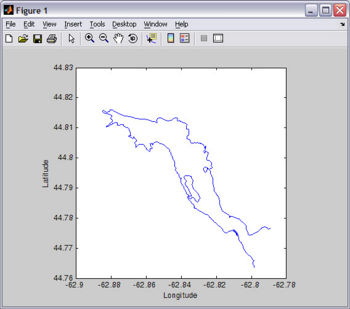
For this tutorial, you will need Fjord_coast.mat. You probably already have this file since it is included in EASYGRID_v0.rar (file from the "Download" section). I created Fjord_coast.mat from the digitized chart I used to make the bathymetry file (see Figure).
![]() If you want to reproduce this plot as an exercise, copy-paste the following code on Matlab's Command Window:
If you want to reproduce this plot as an exercise, copy-paste the following code on Matlab's Command Window:
load Fjord_coast.mat plot(lon,lat); xlabel('Longitude'); ylabel('Latitude'); axis square;
Making your grid
You probably will need to run EASYGRID several times in a iterative manner to create, smooth and mask-edit your grid. You can turn ON or OFF different functionalities of EASYGRID (like the mask-editing and bathymetry-smoothing routines) using the SWITCHES located in the USER SETTINGS section, which is the first part of script (after the help lines). Below a snippet from EASYGRID's code that contains the SWITCHES section.
%********************************************************************************************************
% USER SETTINGS *****************************************************************************************
%********************************************************************************************************
%
% -------------------------------------------------------------------------
% SWITCHES ( ON = 1; OFF = 0 ) --------------------------------------------
% -------------------------------------------------------------------------
create_grid = 1; % Create GRID. Turn OFF to work with a previously created grid (i.e. grid variables
% existing on Workspace)
plot_grid = 1; % Plot grid
smooth_grid = 1; % Smooth bathymetry using H. Arango's smth_bath.m , which applies a Shapiro filter
% to the bathymetry
edit_mask = 0; % Edit rho-mask using interactive plot. Use this to manually change sea-pixels into
% land-pixels and vice-versa
screen_output = 1; % Displays (on screen) many parameters that need to be copy-pasted in the roms.in file
save_grid = 1; % Save GRID in a NetCDF file
save_init = 1; % Create (and save) INITIAL CONDITIONS (from grid)
![]() A typical procedure to create a grid will follow (more or less) these steps:
A typical procedure to create a grid will follow (more or less) these steps:
- TURN ON create_grid and plot_grid (and turn off ALL other switches) and run EASYGRID.
- Tune-up Geographical and Grid parameters and run EASYGRID again.
- Repeat step 2 above until satisfied with the grid location, size and resolution.
- TURN OFF create_grid and proceed to smooth bathymetry by turning ON smooth_grid and re-running EASYGRID.
- Tune-up smoothing parameters and repeat the step above as many times as necessary... when finished, turn OFF smooth_grid.
- Edit land/sea mask by turning ON edit_mask and re-running EASYGRID.
- Tune-up the land/sea mask as many times as necessary... when done, turn OFF edit_mask.
- When satisfied with the grid, bathymetry and mask... save it all to a GRID NetCDF file by turnning ON save_grid and re-running EASYGRID one more time.
Setting up USER SETTINGS
EASYGRID's USER SETTINGS section is where you can change the parameters needed to create your grid and initialization files. To make things easier for you, there are detailed instructions and explanations beside each variable entry line in the USER SETTINGS. In fact, most of the "user-manual" for EASYGRID is included within itself. Therefore I will simply copy-paste below a snippet from EASYGRID's code that contains the USER SETTINGS/instructions section.
![]() NOTE: The default settings in EASYGRID are for the Fjord tidal case, hence you can just run EASYGRID (using the included Fjord_bathy.mat and Fjord_coast.mat files) to generate the grid and initial-conditions files required for the Fjord tidal case tutorial.
NOTE: The default settings in EASYGRID are for the Fjord tidal case, hence you can just run EASYGRID (using the included Fjord_bathy.mat and Fjord_coast.mat files) to generate the grid and initial-conditions files required for the Fjord tidal case tutorial.
%********************************************************************************************************
% USER SETTINGS *****************************************************************************************
%********************************************************************************************************
%
% -------------------------------------------------------------------------
% SWITCHES ( ON = 1; OFF = 0 ) --------------------------------------------
% -------------------------------------------------------------------------
create_grid = 1; % Create GRID. Turn OFF to work with a previously created grid (i.e. grid variables
% existing on Workspace)
plot_grid = 1; % Plot grid
smooth_grid = 1; % Smooth bathymetry using H. Arango's smth_bath.m , which applies a Shapiro filter
% to the bathymetry
edit_mask = 0; % Edit rho-mask using interactive plot. Use this to manually change sea-pixels into
% land-pixels and vice-versa
screen_output = 1; % Displays (on screen) many parameters that need to be copy-pasted in the roms.in file
save_grid = 1; % Save GRID in a NetCDF file
save_init = 1; % Create (and save) INITIAL CONDITIONS (from grid)
% ON = 1; OFF = 0
%
% -------------------------------------------------------------------------
% GRID SETTINGS -----------------------------------------------------------
% -------------------------------------------------------------------------
%
% Geographical and Grid parameters --------
lat = 44.8125; % Latitude (degrees) of the bottom-left corner of the grid.
lon = -62.8855; % Longitude (degrees) of the bottom-left corner of the grid.
%
X = 9100; % Width of domain (meters)
Y = 2550; % Length of domain (meters)
rotangle = -43; % Angle (degrees) to rotate the grid conterclock-wise
resol = 100; % Cell width and height (i.e. Resolution)in meters. Grid cells are
% forced to be (almost) square.
N = 10; % Number of vertical levels
%
%
% Bathymetry -------------- % Bathymetry for ROMS is measured positive downwards (zeros are not allowed)
% see: https://www.myroms.org/wiki/index.php/Grid_Generation#Bathymetry
% To allow variations in surface elevation (eg. tides) while keeping all
% depths positive, an arbitrary offset (see minh below) is added to the
% depth vector.
%
hh = nan; % Analytical Depth (meters) used to create a uniform-depth grid. If using a
% bathymetry file, leave hh = nan;
minh = 4; % Arbitrary depth offset in meters (see above). minh should be a little more
% than the maximum expected tidal variation.
Bathy = 'Fjord_bathy.mat'; % Bathymetry file. If using the analytical depth above (i.e. hh ~= nan), then
% Bathy will not be used. The bathymetry file should be a .mat file containing
% 3 vectors (xbathy, ybathy and zbathy). where xbathy = Longitude,
% ybathy = Latitude and zbathy = depth (positive, in meters). xbathy and ybathy
% are in decimal degrees See: http://en.wikipedia.org/wiki/Decimal_degrees
% Bathymetry smoothing routine... See "Switches section" (above) to turn this ON or OFF
if smooth_grid == 1;
order = 2; % Order of Shapiro filter (2,4,8)... default: 2
rlim = 0.35; % Maximum r-factor allowed (0.35)... default: 0.35
npass = 50; % Maximum number of passes.......... default: 50
end
%---------------------------------
%
%
% Coastline ----------------------
Coast = load('Fjord_coast.mat'); % If there isn't a coastline file... comment-out this line:
% e.g. %Coast = load('Fjord_coast.mat'); The coastline is only used for
% plotting. The coastline .mat file should contain 2 vectors named
% "lat" and "lon"
%
%
% -------------------------------------------------------------------------
% OUTPUT: File naming and NetCDF descriptors ------------------------------
% -------------------------------------------------------------------------
Grid_filename = 'Fjord'; % Filename for grid and initial conditions files (don't include extension).
% "_grd.nc" is added to grid file and "_ini.nc" is added to initial
% conditions file
Descrip_grd = 'Test grid'; %Description for grid .nc file
Descrip_ini = 'Test initial conditions'; %Description for initial conditions .nc file
Author = 'John Smith';
Computer = 'My Computer';
%
% -------------------------------------------------------------------------
% INITIAL CONDITIONS ------------------------------------------------------
% -------------------------------------------------------------------------
if save_init == 1; % See "Switches section" (above) to turn this ON or OFF
% Initial conditions will be constant throughout the grid domain
%--------------------------------------------------------------------------
NH4 = 0.1; % Ammonium concentration (millimole_NH4 meter-3)
NO3 = 10; % Nitrate concentration (millimole_N03 meter-3)
chlorophyll1 = 0.3; % Chlorophyll concentration in small phytoplankyon (milligrams_chlorophyll meter-3)
chlorophyll2 = 0.3; % Chlorophyll concentration in large phytoplankyon (milligrams_chlorophyll meter-3)
detritus1 = 0.03; % Small detritus concentration (millimole_nitrogen meter-3)
detritus2 = 0.03; % Large detritus concentration (millimole_nitrogen meter-3)
detritusC1 = 1; % Small detritus carbon concentration (millimole_carbon meter-3)
detritusC2 = 0.2; % Large detritus carbon concentration (millimole_carbon meter-3)
phyto1 = 0.02; % Small phytoplankton concentration (millimole_nitrogen meter-3)
phyto2 = 0.02; % Large phytoplankton concentration (millimole_nitrogen meter-3)
phytoC1 = 0.2; % Small phytoplankton carbon concentration (millimole_carbon meter-3)
phytoC2 = 0.1; % Small phytoplankton carbon concentration (millimole_carbon meter-3)
salt = 30; % Salinity (PSU)
temp = 9; % Potential temperature (Celsius)
u = 0; % u-momentum component (meter second-1)
ubar = 0; % Vertically integrated u-momentum component (meter second-1)
v = 0; % v-momentum component (meter second-1)
vbar = 0; % Vertically integrated v-momentum component (meter second-1)
zeta = 0; % Free-surface (meters)
zooplankton = 0.01; % Zooplankton concentration "millimole_nitrogen meter-3"
zooplanktonC = 0.5; % Zooplankton carbon concentration "millimole_carbon meter-3"
%--------------------------------------------------------------------------
end
%
%
%********************************************************************************************************
% END OF USER SETTINGS **********************************************************************************
%********************************************************************************************************
![]() NOTE: You may want to change the name of easygrid.m to a name more descriptive of your grid (example: easygrid_fjord.m). That way you can keep many copies of easygrid.m, each with the parameters of each of your different projects.
NOTE: You may want to change the name of easygrid.m to a name more descriptive of your grid (example: easygrid_fjord.m). That way you can keep many copies of easygrid.m, each with the parameters of each of your different projects.
Geographical/Grid parameters
After you got the bathymetry and coastline of your study region, now it is time to place a grid on it... In the USER SETTINGS section, you have to specify lat, lon, X, Y and rotangle. This is done in an iterative manner, where first you input your best-guesses, then you run EASYGRID to plot the grid. From the plot you can adjust your geographical/grid parameters... plot... adjust... plot... adjust... and so on.
![]() You may want to turn off some switches (see below) to speed up the plotting process.
You may want to turn off some switches (see below) to speed up the plotting process.
% -------------------------------------------------------------------------
% SWITCHES ( ON = 1; OFF = 0 ) --------------------------------------------
% -------------------------------------------------------------------------
create_grid = 1; % Create GRID. Turn OFF to work with a previously created grid (i.e. grid variables
% existing on Workspace)
plot_grid = 1; % Plot grid
smooth_grid = 0; % Smooth bathymetry using H. Arango's smth_bath.m , which applies a Shapiro filter
% to the bathymetry
edit_mask = 0; % Edit rho-mask using interactive plot. Use this to manually change sea-pixels into
% land-pixels and vice-versa
screen_output = 0; % Displays (on screen) many parameters that need to be copy-pasted in the roms.in file
save_grid = 0; % Save GRID in a NetCDF file
save_init = 0; % Create (and save) INITIAL CONDITIONS (from grid)
Bathymetry smoothing
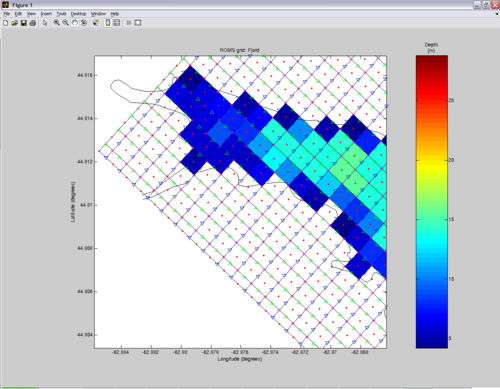
ROMS doesn't like rough bathymetry with abrupt changes in topography. Therefore, after you got your grid where you wanted it to be, the next step is to smooth the bathymetry. Turn ON the smooth_grid variable in the SWITCHES section. Usually simply enabling smoothing (with the default settings) will do a pretty good job. But if unsatisfied, you can iteratively tweak the parameters in the %Bathymetry smoothing section of the USER SETTINGS until obtaining the desired smoothness. Below is a comparison of grids without and with smoothing (default settings).
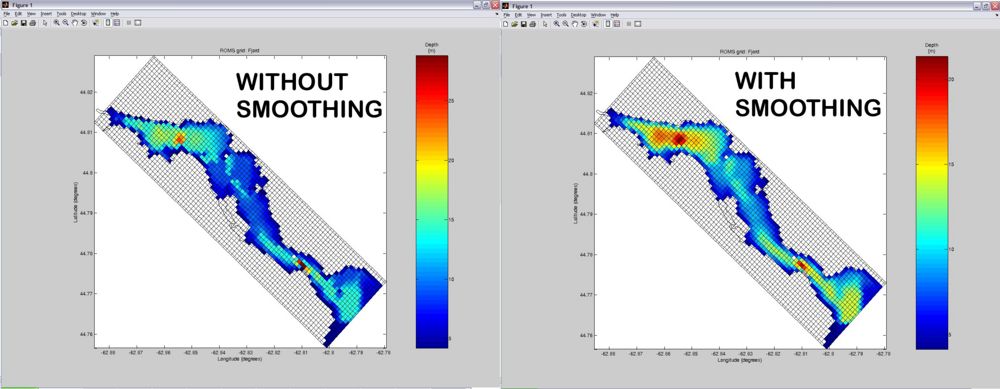
Editing Masks
Since the land/sea mask is automatically created using the bathymetry, it is very likely a few pixels will be wrongly assigned as land or sea. If so, you will need to edit your land/sea mask... and to do this, you can use EASYGRID's Mask-Editing interactive plot, which is a plot of the land/sea mask that lets you (interactive) use your mouse's left-button to toggle land-cells into sea-cells... and vice versa.
First, TURN ON edit_mask and TURN OFF create_grid.
% -------------------------------------------------------------------------
% SWITCHES ( ON = 1; OFF = 0 ) --------------------------------------------
% -------------------------------------------------------------------------
create_grid = 0; % Create GRID. Turn OFF to work with a previously created grid (i.e. grid variables
% existing on Workspace)
plot_grid = 0; % Plot grid
smooth_grid = 0; % Smooth bathymetry using H. Arango's smth_bath.m , which applies a Shapiro filter
% to the bathymetry
edit_mask = 1; % Edit rho-mask using interactive plot. Use this to manually change sea-pixels into
% land-pixels and vice-versa
screen_output = 0; % Displays (on screen) many parameters that need to be copy-pasted in the roms.in file
save_grid = 0; % Save GRID in a NetCDF file
save_init = 0; % Create (and save) INITIAL CONDITIONS (from grid)
Then, run EASYGRID to launch the Mask-Editing interactive plot. You can change sea-cells into land-cells (and vice versa) by clicking on them with your mouse's LEFT-BUTTON. When you are done, simple click on your mouse's RIGHT-BUTTON TO FINISH (i.e. exit toggle mode). If you want to return to toggle mode, simple run EASYGRID again.
Masking Criteria
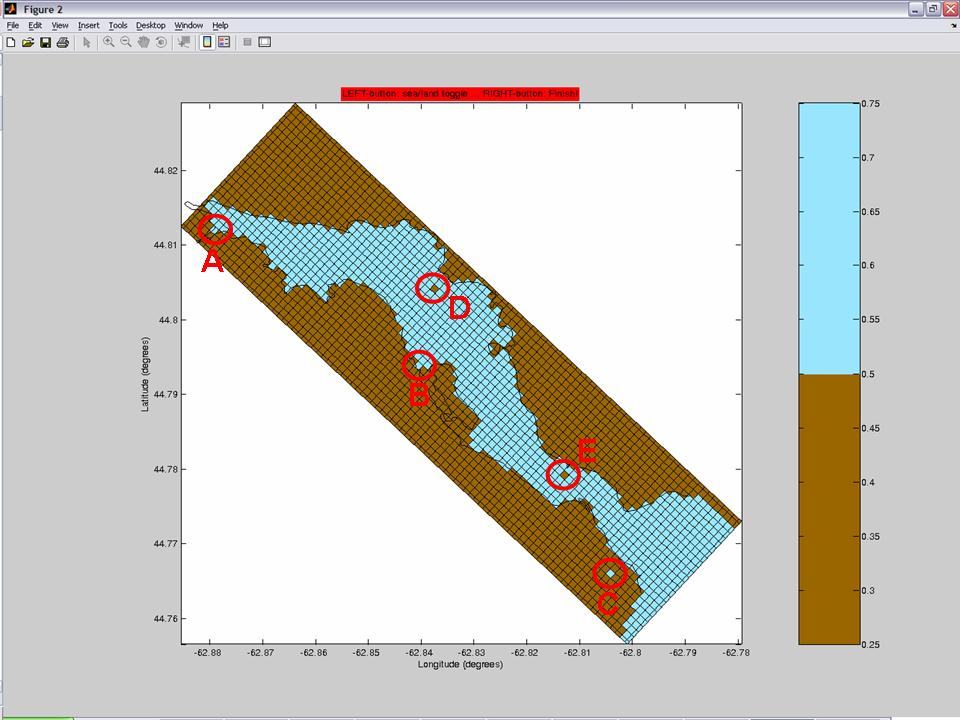
- Mask 1-cell bays (in the figure below, see circles A and B as examples)
- Mask disconnected lakes (in the figure below, see circle C as an example)
- Although tiny islands apparently don't cause troubles... you may want to unmask "islands" that you consider unrealistic or unnecessary (in the figure below, see circles D and E as examples)
Parameter output on screen
If screen_output is turned ON (in the USER SETTINGS section), EASYGRID will output some parameters to screen. Make sure you write this numbers (Lm, MM, N and DT) since you will need to enter them in the roms.in file prior to running ROMS.
| COPY-PASTE the following parameters into your roms.in file | ---------------------------------------------------------------------------------------------- | | | Lm == 90 ! Number of I-direction INTERIOR RHO-points | Mm == 24 ! Number of J-direction INTERIOR RHO-points | N == 10 ! Number of vertical levels | | | Make sure the Baroclinic time-step (DT) in your roms.in file is less than: 11.2939 seconds | ----------------------------------------------------------------------------------------------
- The parameters Lm, Mm and N are the basically the dimensions of the RHO-grid.
- The parameter DT is the baroclinic time-step and it should be less than SQRT( (dx^2+dy^2)/(g*h(x,y) ). EASYGRID solves this equation and tells you the result (11.2939 sec in the case above). You can read a bit more about this in this forum post (middle of page).
Saving GRID to .nc file
Once you got your grid in place, your bathymetry smoothed and your land/sea masks edited... it is time to save your grid in a NetCDF file, which is what ROMS reads. To do this, simply TURN ON save_grid and re-run EASYGRID.
![]() WARNING: You may want to make sure create_grid, edit_mask and smooth_grid are turned OFF since this may ERASE or alter your previous mask-editing or your bathymetry. On the other hand, you may want to make sure screen_output is turned ON so that EASYGRID outputs some parameters to screen (Lm, MM, N and DT), which your should write down, since you will need to enter them in the roms.in file prior to running ROMS.
WARNING: You may want to make sure create_grid, edit_mask and smooth_grid are turned OFF since this may ERASE or alter your previous mask-editing or your bathymetry. On the other hand, you may want to make sure screen_output is turned ON so that EASYGRID outputs some parameters to screen (Lm, MM, N and DT), which your should write down, since you will need to enter them in the roms.in file prior to running ROMS.
Making Initial-Conditions file
ROMS initialization NetCDF files contain information of the spatial distribution of one (or multiple) variable(s) that ROMS uses at the START of the simulation. The value of each initialization variable is specified for each corresponding cell in the ROMS grid... hence, inside a typical initial-conditions file, there will be one three-dimensional matrix (matching grid dimensions) for each specified variable. EASYGRID can generate Initial-Conditions NetCDF files for ROMS, however, only in a rudimentary manner... the user can only choose a single value for each initial-condition variable... EASYGRID makes each variable CONSTANT over the entire grid domain. This may be only useful in some simple cases and to do "quick-tests".
To change the value of a initial-condition variable, simply go to the INITIAL CONDITIONS section in the USER SETTINGS and change the value of the variable of interest.
If you need to add another initial-condition variable... (1) add in the INITIAL CONDITIONS section of the USER SETTINGS:
new_variable = 0; % Variable description (units)
and (2) add the following snippet to the end of the SAVE INITIAL CONDITIONS FILE section of easygrid.m file (substitute the parts in red)
%-------------------------------------------------------------------------- dims = { 'time'; 's_rho'; 'eta_rho'; 'xi_rho'}; nc{ 'newvar'} = ncdouble(dims); nc{ 'newvar'}(:,:,:) = ones(N,Mp,Lp).* 'new_variable'; nc{ 'newvar'}.time = 'ocean_time' ;
![]() WARNING: The dimensions of the new initial-conditions variable should be the same as the grid that will use it. For example, the u momentum initial-condition matrix should have the same dimensions as the u-grid. In a biological setting however, most of the newly-specified variables are likely to be used by the rho-grid.
WARNING: The dimensions of the new initial-conditions variable should be the same as the grid that will use it. For example, the u momentum initial-condition matrix should have the same dimensions as the u-grid. In a biological setting however, most of the newly-specified variables are likely to be used by the rho-grid.
![]() This concludes this tutorial. However, you should consider reviewing the Tidal Fjord TUTORIAL, which uses the grid and initial-condition files generated here to setup a simple tidal realistic application.
This concludes this tutorial. However, you should consider reviewing the Tidal Fjord TUTORIAL, which uses the grid and initial-condition files generated here to setup a simple tidal realistic application.